What do we work on? Like any other structural biology laboratory, we also work primarily on proteins. If you ask what proteins we work on, our answer would be that we work on proteins from viruses, bacteria, yeast, plants, and animals (including humans). Yeah! All of it, actually, and DNA as well. However, it is not the source we are particularly interested in; it is their function that we try to address through structural studies. In our pursuit to ‘solve’ structural problems on a particular theme, we take up proteins from varying sources.
What kind of experiments do we often do? Routine experiments in our laboratory include PCR, cloning, protein over-expression, protein purification, biochemical, biophysical, and functional characterization, crystallization, and structure solution. As a structural biology group, our major tool has been crystallography and, more recently, cryo-EM too! We do quite a lot of SAXS and AUC.
How and where do we get the proteins from? We overexpress most of them in bacteria in recombinant form. We also consider expanding our horizons by expressing proteins in systems like yeast, mammalian, and insect cells.
Do we have a specific area of interest? Over the initial years, we have gained a certain sense of directionality with respect to our science. Details of our current focus areas can be seen below:
(1) Chromatin Structural Biology: Histone proteins package eukaryotic DNA into chromatin. The dynamic DNA-protein complex called nucleosomes forms the basic element of chromatin. All eukaryotic genomic processes, both natural and pathological,  including transcription, replication, recombination, repair, chromosome segregation, carcinogenesis, and viral infection, can be fully understood only in the context of nucleosomes. The core of a nucleosome, called the nucleosome core particle (NCP), consists of fourteen turns of B-form DNA around an octamer of histone proteins, as can be seen in the image here.
including transcription, replication, recombination, repair, chromosome segregation, carcinogenesis, and viral infection, can be fully understood only in the context of nucleosomes. The core of a nucleosome, called the nucleosome core particle (NCP), consists of fourteen turns of B-form DNA around an octamer of histone proteins, as can be seen in the image here.
A nucleosome is known to be the binding platform for several protein factors. For a protein to gain access to chromosomal DNA/histone proteins, it needs to interact and/or compete with the histone proteins/DNA of the nucleosome. Studies involving nucleosomes are of greater significance than isolated studies focusing on the interaction of protein factors with individual histone(s) or DNA. In this context, we work on some nucleosome-interacting proteins.
Around the theme of chromatin structural biology, another of our interests is histone chaperones. Histone chaperones are a group of proteins that bind histones and regulate nucleosome assembly. The mode of action of several histone chaperones is not well understood. Also, it is unclear what decides the specificity of certain histone chaperones for H3/H4, H2A/H2B, or histone variants. To better understand these molecular machines, we work on a few projects dealing with the structural characterization of some important but poorly characterized histone chaperones. A major interest in this direction has been nucleoplasmins.
(2) Structural Biology of Clp Machinery Proteins: The caseinolytic protease (Clp) machinery is present in bacteria, plants, and other eukaryotes. These proteases usually 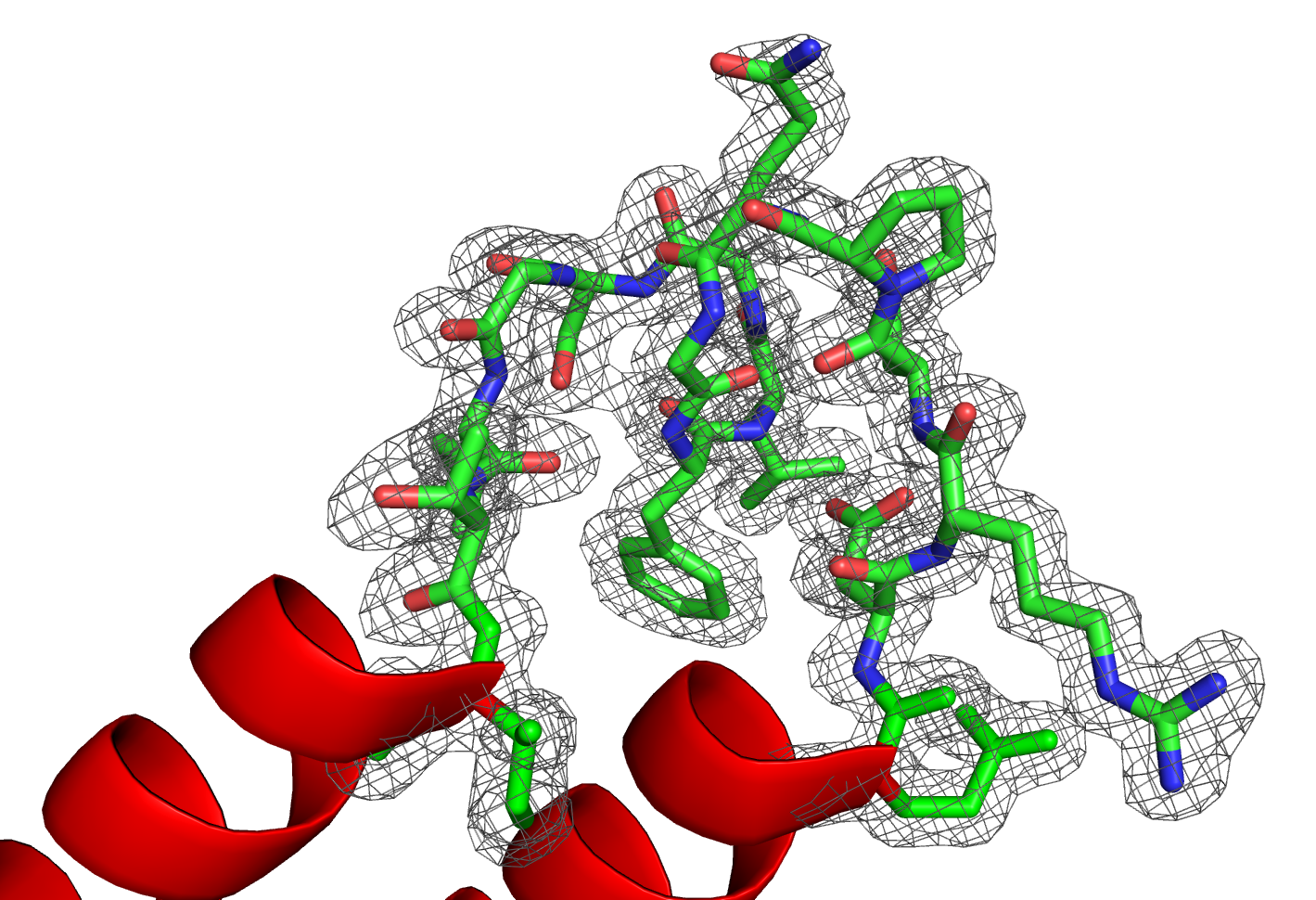 associate with Clp chaperone proteins that help in the ATP-dependent unfolding of substrates, which need to be taken in for proteolysis by the proteases. ClpD is a Clp-associated chaperone unique to plants. Plant ClpC and ClpD proteins both get localized in the chloroplast stroma. However, our work from ILS has shown that the structure of ClpD is quite different from that of ClpC in its N-terminus. We are interested in plant Clp machinery proteins to understand their distinct roles in protein homeostasis and stress response. Additionally, we work on mycobacterial Clp chaperones due to their importance as a target in anti-mycobacterial research. A project on this theme has been on the anti-mycobacterial peptide Lassomycin, which targets MtClpC1.
associate with Clp chaperone proteins that help in the ATP-dependent unfolding of substrates, which need to be taken in for proteolysis by the proteases. ClpD is a Clp-associated chaperone unique to plants. Plant ClpC and ClpD proteins both get localized in the chloroplast stroma. However, our work from ILS has shown that the structure of ClpD is quite different from that of ClpC in its N-terminus. We are interested in plant Clp machinery proteins to understand their distinct roles in protein homeostasis and stress response. Additionally, we work on mycobacterial Clp chaperones due to their importance as a target in anti-mycobacterial research. A project on this theme has been on the anti-mycobacterial peptide Lassomycin, which targets MtClpC1.
(3) Infectious Disease Structural Biology: A couple of our chromatin structural biology and Clp machinery projects have an infectious disease flavor. We have ongoing projects on proteins from bacteria, fungi, viruses, and parasites that infect humans. Moreover, we work on candidate proteins from pathogens that infect cultured shrimps. We are initiating a project on the structural and functional characterization of certain antimicrobial peptides that target bacterial membranes.
Over and above these, additional crystallographic projects are being taken up collaboratively from time to time.
Whom do we collaborate with? We do have several collaborators.
Our active collaborators from ILS are:
- Dr. Balachandran Ravindran (immunology)
- Dr. Narottam Acharya (yeast genome instability)
- Dr. Tushar Kant Beuria (bacterial cell division)
Our active collaborators from RGCB are:
- Dr. Kathiresan Natarajan (neurotransmitter receptors)
- Dr. Lightson N.G. (aptamer-protein complexes)
- Dr. Mahendran K.R. (transmembrane pores)
- Dr. Rajesh Chandramohanadas (red blood cell pathology)
In addition, we have ongoing collaborative projects with the following:
- Prof. Aline V. Probst, CNRS, France (histone variants and chaperones)
- Dr. Devinder Sehgal, NII, New Delhi, India (bacterial virulence)
- Prof. Klaas van Wijk, Cornell University, USA (plant protein homeostasis)
- Dr. Sandip Basak, NTU, Singapore (cryo-electron microscopy)
Are we keen on new collaborations? As a structural biology laboratory, we believe in evolving by adding on newer projects and starting new collaborations. Each project, in fact, comes with a good learning experience for our lab members. Our hands are relatively full at the moment. Due to the inherent risk involved in structural biology projects, each of us juggles with 3-4 projects at any given time, 1-2 projects of the lab’s main theme, and others, mostly collaborative! Having said this, you could still persuade us to pursue a bit of your science. We don’t say ‘NO’ that easily. 🙂